t-SNE
t-SNE (t-distributed Stochastic Neighbourhood Embedding) is a nice statistical method that can be used to visualise high-dimensional data in a smaller number of dimensions.
I guess I could say I learnt about this method in the context of my MSc thesis, and even though I only used it once yet, I found the results to be absolutely great!
I don't want to give too much detail (and I don't think I could, if I wanted...) but I was working with a space of 24 dimensions. (If there are people who struggle with doing 3D visualisation in their heads, imagine 24D!) Despite the fact that the data had 24 dimensions, I knew (from the context) that there were two main groups of datapoints, with one of them accounting for roughly 25% of the data.
I applied t-SNE to that data, roughly 10.000 datapoints of that 24D space, and the results I got looked really great, to be honest. Below, you can see the result I got with minimal effort, and you can even see that it does a pretty decent job of separating the two groups of data:
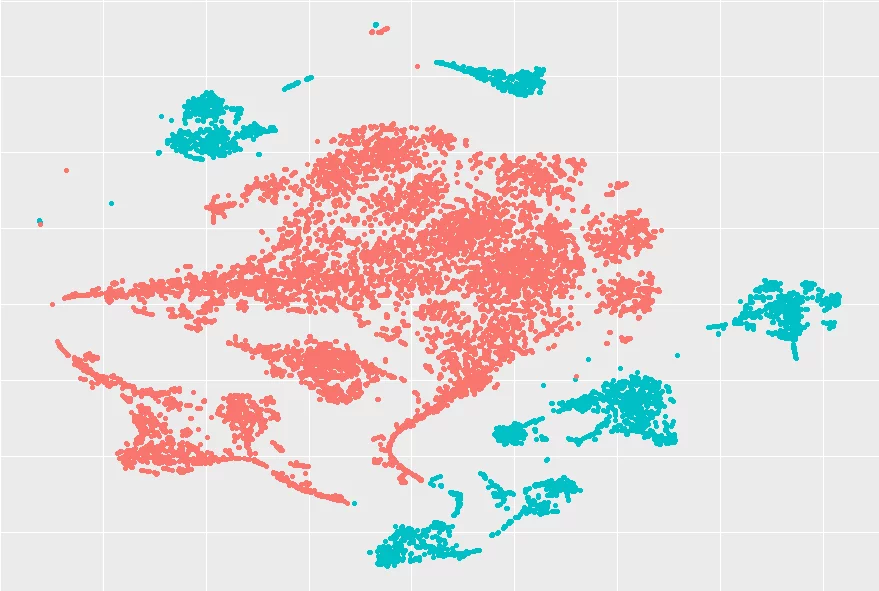
I was really impressed by the results, and I was also very curious about how hard it would be to implement this method. From the paper referenced below, the method doesn't seem hard to implement, so I might give it a go...
(By the way, if you are into Python, you can use t-SNE with the package sklearn.
That's it for now! Stay tuned and I'll see you around!
Become the smartest Python 🐍 developer in the room 🚀
Every Monday, you'll get a Python deep dive that unpacks a topic with analogies, diagrams, and code examples so you can write clearer, faster, and more idiomatic code.
References
- Van der Maaten, L. and Hinton, G., 2008. Visualizing data using t-SNE. Journal of machine learning research, 9(11).
- t-SNE, Wikipedia, https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding [last accessed 19-02-2022];
